Table of Contents
Citracer
📝 Description
Trace citation chains for any keyword across research papers.
Given a source PDF and a keyword, citracer parses the bibliography with GROBID, finds every occurrence of the keyword in the body, identifies the references cited near each occurrence, downloads those papers, and recursively walks the resulting citation graph. The output is an interactive HTML page.
Supported sources. citracer currently resolves cited papers through three external services: arXiv, Semantic Scholar, and OpenReview (for ICLR / TMLR papers not on arXiv). Workshop proceedings, books, and paywalled journal articles are not retrievable and appear as
unavailablenodes in the graph.
⚙️ Installation
Requirements: Python 3.10+ and Docker.
GROBID must be reachable on http://localhost:8070. Verify with curl http://localhost:8070/api/isalive.
A Semantic Scholar API key is optional but recommended — without one the public endpoint is throttled to ~3.5s between calls. With a key, the throttle drops to 0.2s.
🚀 Usage
Flag | Default | Description |
|---|---|---|
| required | Path to the source PDF |
| required | Term to trace through citations |
|
| Maximum recursion depth |
| off | Show passages directly in node tooltips |
|
| Output HTML file |
|
| Local cache for PDFs and metadata |
|
| GROBID service URL |
| none | Semantic Scholar API key |
| sentence-based | If set, fall back to a ±N character window for ref association |
| off | Do not open the result in a browser |
| off | Verbose logging |
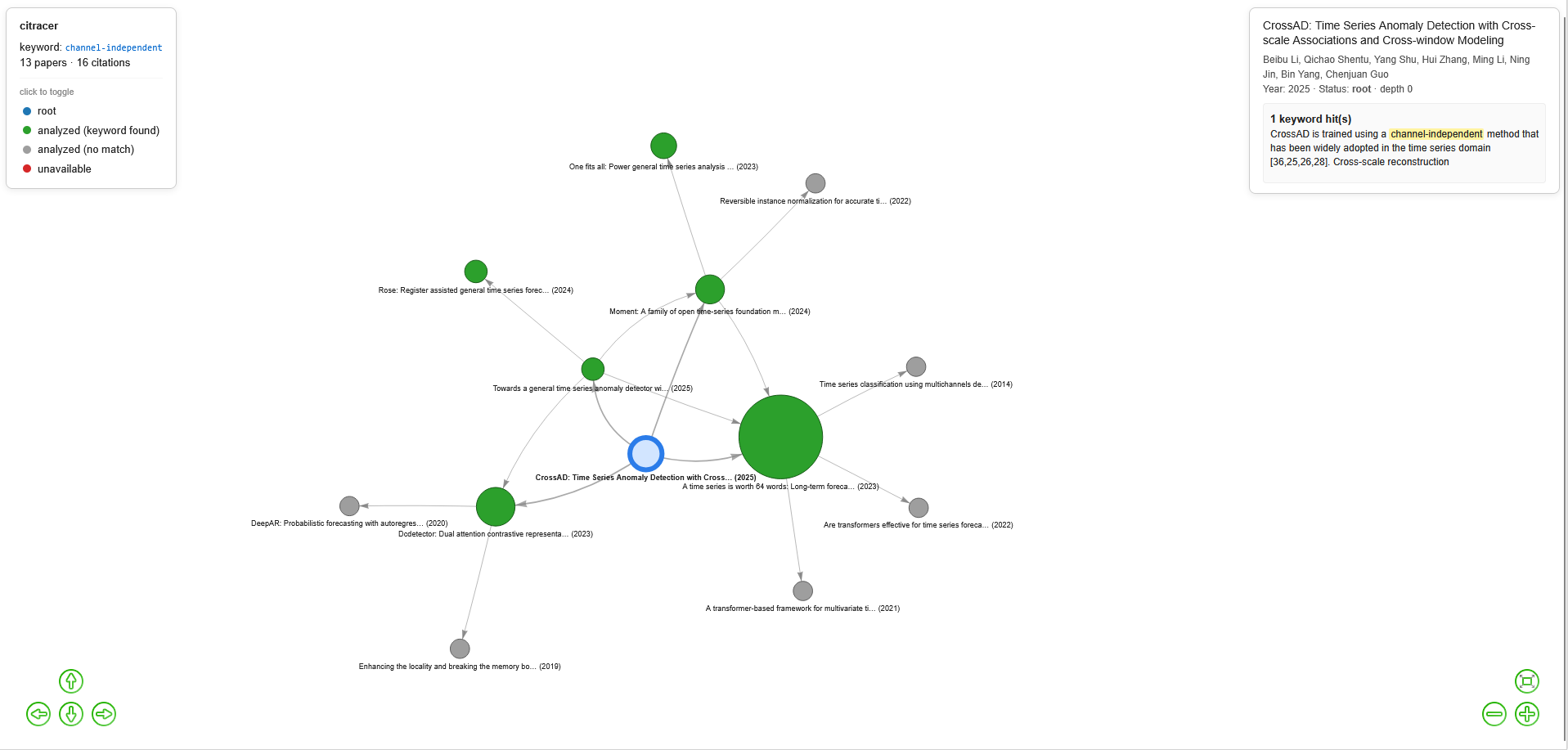
🎨 Output
Nodes are colored by status:
Color | Status | Meaning |
|---|---|---|
blue |
| The source PDF |
green |
| PDF retrieved and the keyword was found in its text |
gray |
| PDF retrieved and parsed, but the keyword does not appear |
red |
| PDF could not be retrieved |
Node size scales with the number of keyword occurrences. The interactive graph supports hover for live preview, click to pin a node, click on the legend to toggle visibility by status, and KaTeX rendering of LaTeX in passages.
🔍 How it works
PDF parsing. GROBID processes the PDF and returns TEI XML. citracer walks the
<body>to reconstruct the plain text while recording the character offset of every inline<ref type="bibr">citation. The bibliography is extracted from<listBibl>. Figure-diagram paragraphs (detected by their density of mathematical Unicode characters) are skipped to avoid polluting the keyword matcher.Keyword matching. The keyword is compiled to a flexible regex that handles morphological variants (e.g.
channel-independentmatcheschannel-independence,channel independently,channelindependence). The body is segmented into sentences with pysbd, and each occurrence of the keyword is associated with the references cited in the same sentence or the immediately following one.Reference resolution. Each cited paper is resolved through the following cascade:
- If GROBID extracted a DOI or arXiv ID, use it directly.
- Otherwise, search arXiv by title (phrase first, then keyword fallback, with rapidfuzz validation).
- If arXiv has nothing, query Semantic Scholar with 429-aware backoff.
- As a last resort, search OpenReview (covers ICLR/TMLR papers not on arXiv).
Resolved PDFs are cached in
./cache/pdfs/.Recursion. The tracer is a BFS that processes papers in queue order, deduplicating by canonical ID (DOI > arXiv > OpenReview > title hash). When the same PDF is reached via a second path, the new edge is added without re-parsing.
Rendering. The graph is serialized to an interactive HTML page using pyvis, with a custom overlay for the legend filter, side info panel, keyword highlighting, and KaTeX math.
📁 Project structure
citation_tracer/
├── cli.py # argparse entry point
├── pdf_parser.py # GROBID + TEI walking + figure-noise filter + pymupdf fallback
├── keyword_matcher.py # morphological regex + sentence-based ref association
├── reference_resolver.py # arXiv-first cascade resolver with cache
├── tracer.py # BFS recursion with deduplication
├── visualizer.py # pyvis rendering + custom overlay
├── models.py # dataclasses
└── utils.py # ID normalization, hashing, logging
🧩 Dependencies
Package | Used for |
|---|---|
PDF structural parsing (external service) | |
TEI XML processing | |
PDF text extraction (parser fallback) | |
arXiv search and download | |
Sentence boundary detection | |
Interactive HTML graph rendering | |
Fuzzy title matching | |
HTTP client | |
Progress bar | |
LaTeX math rendering in the HTML output (CDN) |
External APIs:
⚠️ Limitations
- GROBID misclassifies a small fraction of references (especially sub-citations like
Liu et al., 2024b). These are silently dropped. - pysbd handles most academic abbreviations but can occasionally split mid-sentence; falling back to
--context-window 300is sometimes useful. - arXiv enforces ~3 seconds between requests, so the first run on a deep trace can take several minutes. The local cache makes subsequent runs fast.
- Only three sources are supported for resolving cited papers: arXiv, Semantic Scholar and OpenReview. Workshop papers, books, and journal articles without an open-access PDF on one of these platforms appear as
unavailablered nodes.
✍️ Authors
- Marc Pinet - Initial work - marcpinet