citracer
A paper cites 50+ references, but which ones actually discuss the concept you care about? And which papers those cite? And the ones after that?
citracer answers this recursively. Give it a PDF and a keyword: it finds every sentence where the keyword appears, identifies the references cited nearby, downloads those papers, and repeats the process N levels deep. A 5-depth trace starting from a single paper typically surfaces 50-150 relevant papers in minutes.
With --semantic, matching goes beyond literal keywords: a sentence-transformer model catches passages that express the same concept with different vocabulary.
Get started in 30 seconds
pip install citracer
docker run --rm -p 8070:8070 lfoppiano/grobid:0.9.0
citracer --pdf paper.pdf --keyword "channel-independent" --depth 3
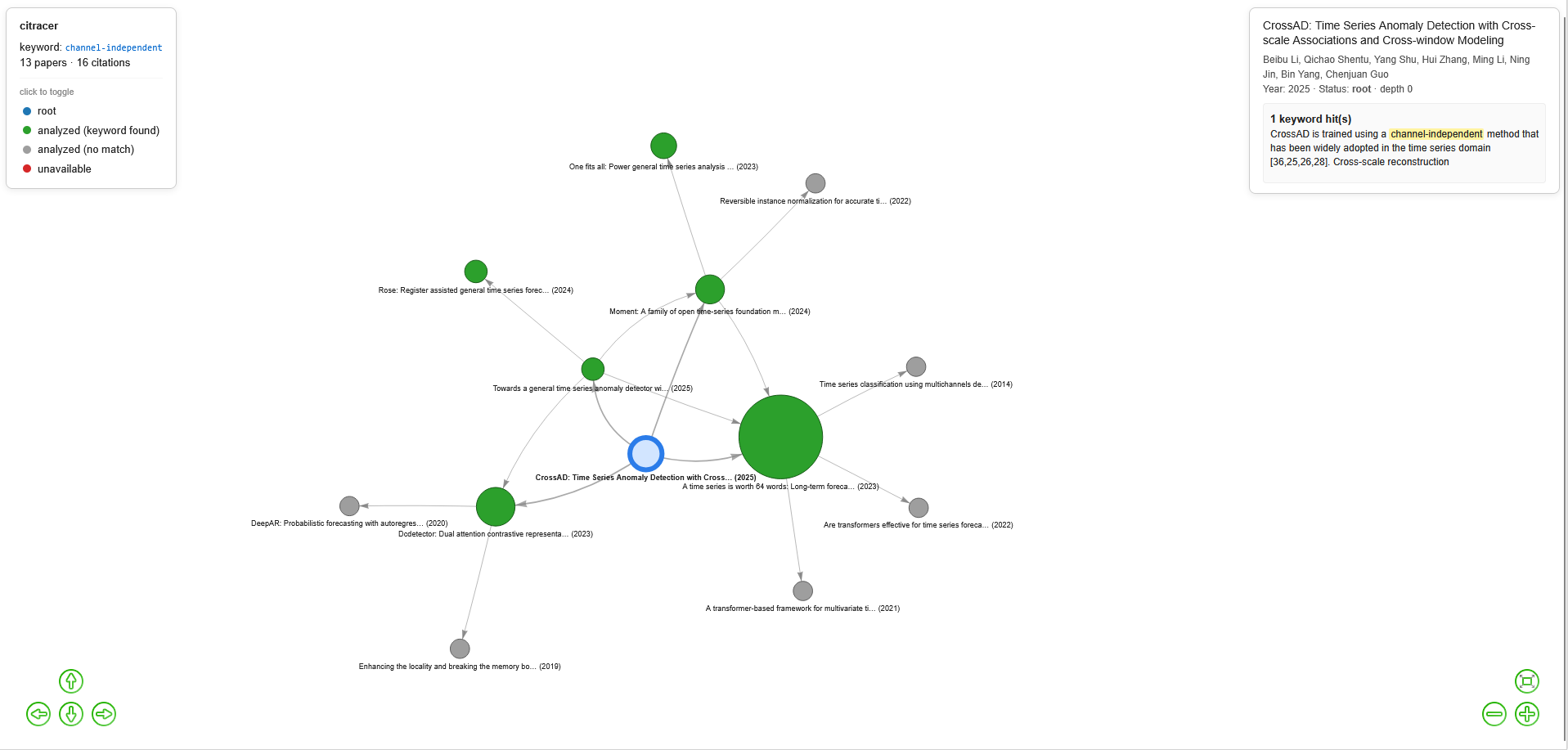
This parses the PDF, finds sentences mentioning your keyword, downloads the papers cited nearby, repeats for 3 levels, and opens an interactive graph in your browser.
Key features
- Recursive tracing: walk N levels deep through citation trees, keeping only papers that discuss your concept
- Concept matching (
--semantic): go beyond literal keywords with sentence-transformer embeddings - Reverse trace (
--reverse): find papers that cite a source while mentioning the keyword - 10+ paper sources: arXiv, Semantic Scholar, OpenReview, Sci-Hub, bioRxiv, medRxiv, ChemRxiv, SSRN, PsyArXiv, AgriXiv, engrXiv
- Literature monitoring (
--diff): compare traces over time and highlight new papers - Interactive HTML output with layout controls, search, filtering, and analytics
- Bibliometric analytics: PageRank, betweenness centrality, pivot detection, keyword density timeline
- Export to JSON and GraphML for downstream analysis
- Reproducibility manifest encoding all trace parameters